As part of their PhD research projects within TransPot and proEVLifeCycle networks, Annelies Van Hemelryk (Erasmus MC) and Ingrid Tomljanovic (GenomeScan) recently established and characterized a novel panel of preclinical models of castration-resistant prostate cancer. These models are derived from tumor samples collected from patients who received a diversity of modern therapies. Through histology, genotyping, and gene expression analyses, this work showed that the models not only retain original tumor characteristics, but also respond to treatments in a way that matches the original patient responses. As a result, they provide new venues for personalized medicine development in advanced prostate cancer.
Prostate cancer is the second most frequent cancer in men worldwide. Castration-resistant prostate cancer (CRPC) is an advanced and aggressive form of the disease which remains incurable and lethal. There is an urgent need for developing personalized medicine approaches that could lead to a better diagnosis, prognosis, and treatment of CRPC patients. Therefore, it is crucial to generate preclinical models of prostate cancer that adequately represent patient tumors and as such can be used to advance translational research.
The diversity of modern treatment options in CRPC has improved patient outcomes, while also likely contributing to an increase in disease heterogeneity. Consequently, there is also a lack of preclinical models that represent the current clinical landscape of CRPC.
Models represent original tumor characteristics and drug responses
To address the need for models that represent tumor heterogeneity and differential responses to treatment, new preclinical models including organoid cultures were developed in the Experimental Urology Department of Erasmus MC. Morphological characterization was complemented by genomic and transcriptomic analyses performed by GenomeScan. Together, these assessments confirmed model consistency between patient materials and preclinical models.
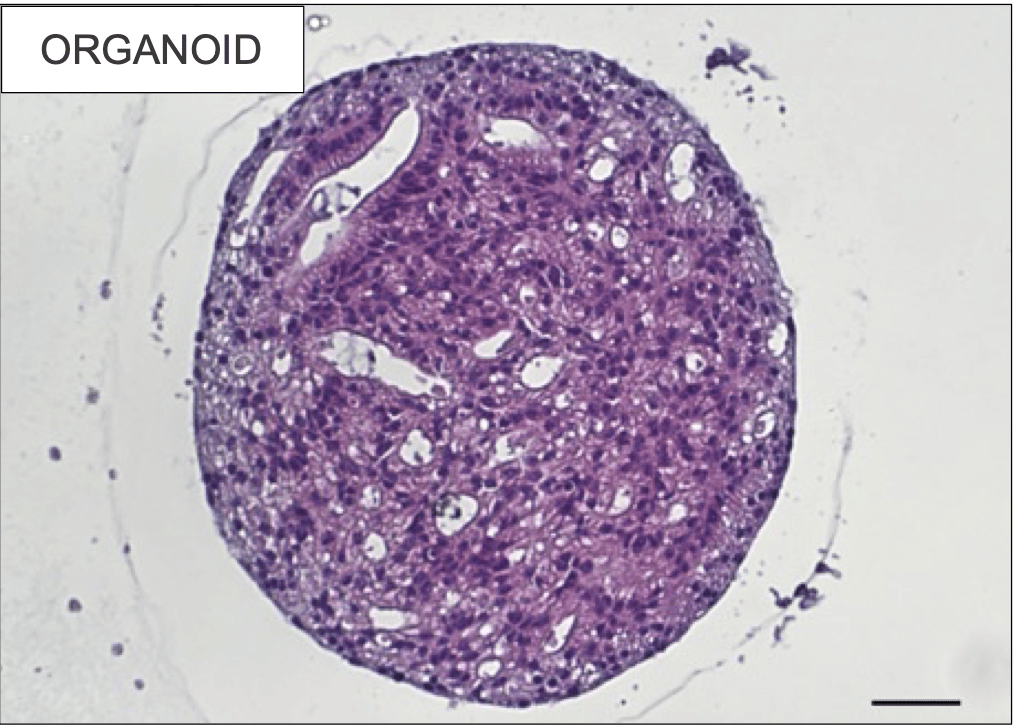
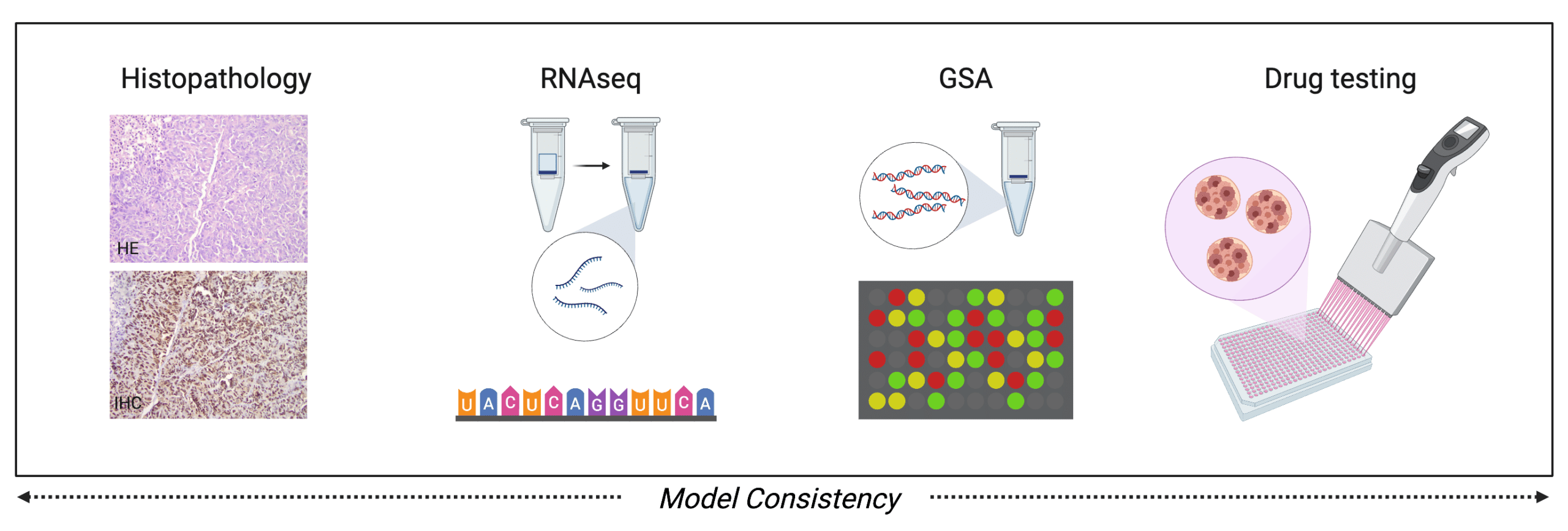
Tumor samples and the preclinical models were all thoroughly characterized by morphological analyses, as well as SNP genotyping arrays and RNA sequencing, confirming model consistency. The preclinical models recapitulated matching patient drug responses, making them a valuable drug testing platform for personalized therapies against castration-resistant prostate cancer.
The genomic and transcriptomic analyses also provided in-depth insights about the models in comparison to large CRPC patient cohorts. Genomic alterations commonly found in prostate cancer were identified. These included copy number alterations in several genes of the androgen receptor pathway, which is known to play a prominent role in prostate cancer. Additionally, the new preclinical models harbored a range of genomic alterations in members of other clinically actionable pathways, which opens opportunities for drug testing.
Importantly – and rather uniquely for prostate cancer research – drug responses in the models revealed a close correspondence to original patient responses.
Significance and collaboration
In summary, this work spans the establishment and in-depth characterization of patient-derived preclinical models that faithfully represent castration-resistant prostate cancer and thereby constitute a valuable resource to study underlying disease mechanisms. The significance of establishing new organoid cultures is underscored by their high level of scalability, which allows them to be leveraged for high-throughput research studies and exchange across the research community. As such, they carry broader relevance in the context of accelerating future efforts in prostate cancer personalized medicine.
This work was performed as a collaboration between GenomeScan and Erasmus MC within the scope of TransPot and proEVLifeCycle research networks. These networks consist of institutions across Europe and share a focus on tackling prostate cancer with innovative and interdisciplinary approaches. The research was funded by the European Union’s Horizon 2020 research and innovation program under the Marie Skłodowska-Curie grant agreements No 721746 and No 860303.
Further reading
Read the full manuscript here.
Learn more about proEVLifeCycle and TransPot, and explore other projects in our portfolio.





