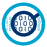
The amplification of the 16S prokaryote-specific gene allows you to analyze the microbiome together with the host without any loss of efficiency or sequencing depth. GenomeScan’s scientists have designed an optimized library preparation method, leading to a higher yield than other kits available on the market. A dual-indexing strategy ensures that hundreds of samples can be analyzed together, making this a very reliable and powerful tool.

16S microbial profiling
Why use 16S profiling?

How 16S profiling works
Identification and comparison of the microbes present in your samples is performed by amplification of the fourth hypervariable domain (V4) of the 16S ribosomal RNA gene. The 16S region is a conserved region of the prokaryotic ribosome but it also contains hypervariable sequences that provide the species-specific signature.
- Taxonomic classification in complex mixtures
- Detailed overview of the prokaryotic composition to family, genus and sometimes species level
- Relative frequencies in comprehensive report
- Publication-ready figures
Data analysis
GenomeScan BioIT experts have set up a data-analysis pipeline to efficiently process all microbiome data. Sequential steps include data trimming and preparation for alignment to the NCBI reference database to determine the characteristics and relative abundancies of different microbial populations.
The output consists of a comprehensive table containing the various microbes that were detected. It displays the taxonomic rank, genome size and number of reads that classifies the various microorganisms. Krona-plots provide an interactive tool to visualize the composition of the sample.

We have summarized key information about our 16S microbial profiling service into a service specification sheet.
Let's get the conversation started for your next NGS project
Please either fill in below or email us directly at info@genomescan.nl
and we will get in touch with you to discuss your requirements





